|
 |
- Search
| Psychiatry Investig > Volume 14(5); 2017 > Article |
Abstract
Objective
Circadian rhythms have been linked to psychiatric disorders such as Depression and Bipolar Disorder (BD). Given previous evidences of sleep/circadian disturbances as well as the genetic susceptibility for BD, we decided to investigate the possible link between the PERIOD3 (Per3) circadian gene and BD.
Methods
This is a genetic association case (BD) vs. control study of the Per3 gene. We further subdivided our BD sample into “good sleepers” (PSQI ≤5) and “poor sleepers” (PSQI>5) according to the Pittsburgh Sleep Quality Index (PSQI) global score, and then we assessed genetic association of the Per3 gene with sleep quality in the BD group.
Results
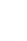
There were 209 cases and 213 controls in our sample. The GT genotype of the SNP rs707467 significantly associated with BD (χ2=8.80; p-value=0.01; adjusted residual=±2.6). We also found significant association of the SNP rs10462020 allele T with BD (χ2=5.81; p-value=0.01) as well as the genotype TT (χ2= 6.01; p-value=0.04; adjusted residual=±2.4).
Circadian rhythms are defined as bioquemical, physiological and behavioral variables, which show an approximately 24-hour cycle oscillation, reacting primarily to the light-dark environmental cycle.1 Preceding studies indicate they are found throughout wide-ranging species, including animals, plants and microorganisms.23 Chronobiology studies biological rhythms, of which circadian rhythms are an example.4
The central biological “clock” that controls circadian rhythms is located in the hypothalamus, consisting of a group of nearly 20,000 neurons named suprachiasmatic nucleus (SCN).2 Previous studies have already recognized genes that drive circadian rhythms in humans and numerous other species, which are frequently called circadian or “clock” genes.56 This group of genes are highly expressed in the SCN and their coordinated operation is responsible for the translation of the environmental circadian input into the organism.567
Circadian rhythms can influence sleep-wake cycles, hormone release, body temperature and other important bodily functions.3 They have been linked to various sleep disorders.8 Abnormal circadian rhythms have also been associated with obesity, diabetes, and psychiatric disorders such as Depression and Bipolar Disorder (BD).910
Some prior studies have revealed that, along with features connected to the sleep-wake cycle (chronotype, actigraphic measures), other variables related to circadian rhythms appear to be altered in BD. In fact, abnormalities of circadian rhythms such as body temperature, blood pressure, heart rate, hormone and neurotransmitters levels have been identified in patients with BD.1112 Circadian rhythms are important in determining human sleep patterns as well.13 Likewise, researchers have demonstrated that sleep disturbances are frequent between BD patients.14 Moreover, we have recently presented that poor sleep quality is a prevalent feature in BD patients even during the interepisodic phases of the disorder.15
Bipolar Disorder is a complex and multifactorial psychiatric disorder.16 Previous evidences from genetic research have confirmed that BD is highly heritable.17 Even though BD exhibits significant genetic vulnerability, the specific underlying genetic factors remains unclear.17 Growing amounts of evidences have implicated circadian genes in the heritability of BD. A number of genetic association studies have demonstrated associations between BD and various circadian genes, such as the Clock, Npas2, Artnl1, Per3 and Nr1d1 genes.18
The human PERIOD3 (Per3) gene is an important member of the Period family that was first isolated in mammals in 1998.19 It is located on human chromosome 1 (1p36.23) and is highly expressed in the suprachiasmatic nucleus.19 One study have recently showed that lithium, one of the first choice pharmacological agents for BD, alters the expression of the Per3 gene in laboratory cultures of fibroblasts.20 Genetic linkage studies of BD showed positive results for the chromosome 1 region where the Per3 gene is located.2122
Genetic association studies have shown some positive results of the Per3 gene and BD plus certain clinical subphenotypes, although there have been preceding publications that found no evidence of association between the Per3 gene and BD.2324252627282930 Thus, given the previous evidences of sleep/circadian disturbances as well as the genetic susceptibility for BD, we decided to investigate the possible link between the Per3 circadian gene and BD. The aim of the present study is to assess the association of some polymorphisms of the Per3 gene in a sample of BD patients.
This report consists of a genetic association case (BD) vs. control study of the Per3 gene. Data collection took place from 2009 to 2013. There were 209 cases and 213 controls in our sample. We further subdivided our BD sample into “good sleepers” (PSQI ≤5) and “poor sleepers” (PSQI>5) according to the Pittsburgh Sleep Quality Index (PSQI) global score, and then we assessed genetic association of the Per3 gene with sleep quality in the BD group (cases).31
The PSQI is a subjective sleep quality instrument, widely used in clinical and research settings and validated to the Brazilian Portuguese.32 The instrument contains seven sub-components, which are relevant features of sleep quality, and generates a global score ranging from 0-21. Global PSQI scores >5 indicates poor sleep quality, whereas scores ≤5 suggests good sleep quality. In order to keep mood variations impact on sleep patterns in a minimum level, we picked BD cases fulfilling criteria for euthymia or remission.33 Euthymia was settled as a score lower than seven in both Hamilton and Young mood scales for the last month along with the lack of criteria for current depressive, mixed or hypo/manic episode by the DSM-IV-TR criteria.333435 The MINI-plus structured interview based on DSM-IV diagnostic criteria was used to set up for psychiatric diagnosis in the BD group.36 Psychiatric longitudinal follow-up evaluated by medical records were also taken into account to endorse accuracy of psychiatric diagnosis. Our sample of cases was composed by adult BD outpatients of Hospital das Clínicas-UFMG psychiatric service.
Controls were recruited from the general population. Inclusion criteria for the control group were an age ≥18 and the lack of criteria for psychiatric diagnosis by the MINI-plus, according to the DSM-IV.
We collected peripheral blood samples from cases and control groups. Genomic DNA was isolated with the saline method.37 DNA quantification was performed on spectrophotometer device, and subsequently, its concentration was adjusted to 50 ng/µL. Genotyping was achieved by Real-Time PCR (RT-PCR) and allelic discrimination was completed by Applied Biosystems (ABI) producer's protocol in the 7500 RTPCR System. The protocol used TaqMan® Genotyping Master Mix (Applied Biosystems, Foster City, CA, USA).
The Taqman® is a type of assay supplied by ABI (Applied Biosystems, Foster City, CA, USA) in the versions assay-ondemand and assay-by-design, consisting of two pairs of oligonucleotide primers: forward and reverse (oligonucleotide concentration in 900 µM), a marker dye VIC®-detects the presence of the allele 1 (markers with final concentration 200 µM), a second marker FAMTM dye-detects the presence of allele 2 (markers with final concentration of 200 µM). PCR reaction included an initial denaturation at 95℃ for 10 min followed by 50 cycles at 95℃ for 15 sec and 60℃ for 1 min. Every reaction contained 5.0 µL of mix, 0.1 µL of probe, 3.9 µL of deionized water and 60 ng of DNA. Ten percent of the samples were genotyped again at random, as a quality control approach, to ensure the exclusion of possible laboratory errors.
We picked single nucleotide polymorphisms (tagSNP's) that would provide a good cover of the whole Per3 gene using HapMap database (www.hapmap.org) and prior published studies. We further used 20 ng of DNA of each patient for five TaqMan SNP's genotyping assays (rs228697, rs228727, rs228729, rs707467, and rs10462020) according to the tutoring of Applied Biosystems.
Gender, genotype and allelic distributions were analyzed by chi-square or exact Fisher test using the software SPSS 20.0. Age was analysed by Mann Whitney non-parametric test. We took p-values of 0.05 or less as evidence of statistical significance. Adjusted residuals, which enables us to analyse the residuals in each cell of the cross tabulation, were performed using a cut-off point of +1.96 and -1.96.
There were 209 cases and 213 controls in our sample. Groups were similar regarding age (Mann-Whitney U=22096.50; p-value=0.897) and gender (χ2=1.66; p-value=0.197). Median age were 43 for cases (min 18; max 88) and 42 for controls (min 18; max 86). 31.60% of cases were males, while 37.60% of controls were males. The SNP rs228697 failed to reach Hardy Weinberg equilibrium (HWE) and it was excluded of further analysis (χ2=17.07; p-value<0.01). All other SNP's were in HWE, except for rs10462020 which violated HWE as regards to the whole sample (χ2=7.20; p-value<0.01), although the control group was in HWE (χ2=1.92; p-value=0.16).
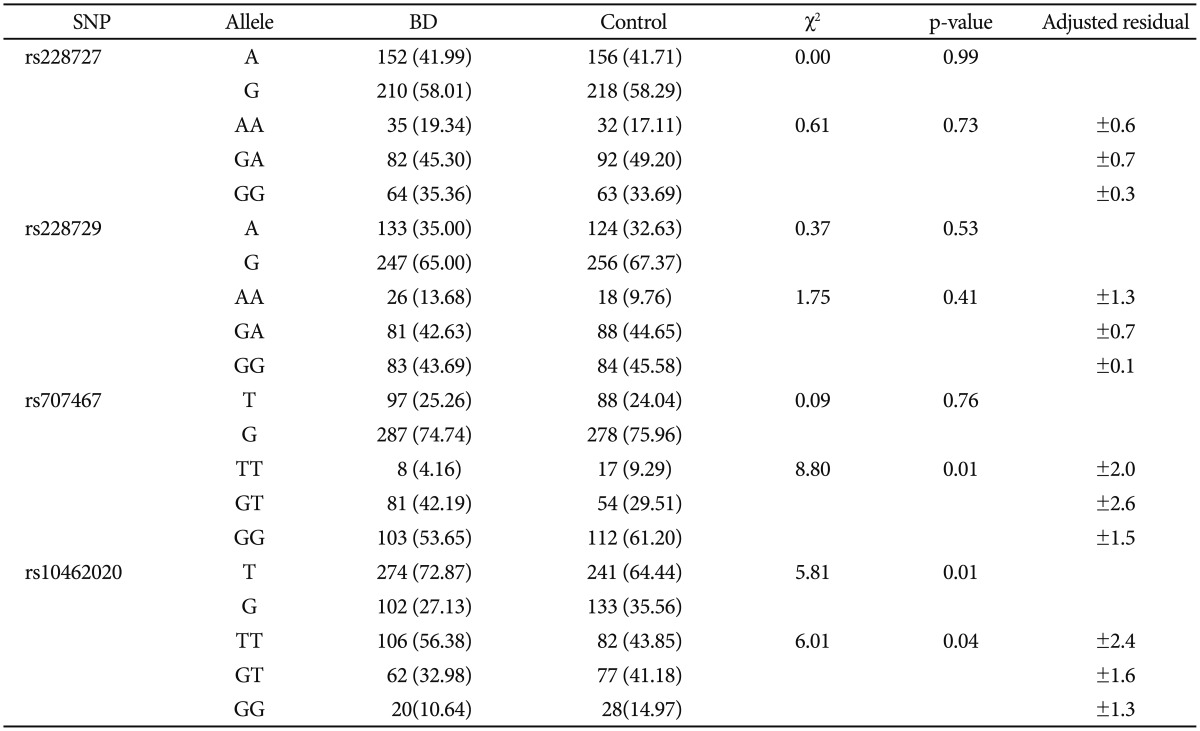
Table 1 shows the genotypic and allelic distributions of the Per3 gene SNP's assessed between cases and controls. The GT genotype of the SNP rs707467 significantly associated with BD (χ2=8.80; p-value=0.01; adjusted residual=±2.6). We also found significant association of the SNP rs10462020 allele T with BD (χ2=5.81; p-value=0.01) as well as the genotype TT with BD (χ2=6.01; p-value=0.04; adjusted residual=±2.4).
Table 2 shows the genotypic and allelic distributions of the Per3 gene SNP's assessed in the BD group subdivided into “good” and “poor” sleepers measured by the global PSQI score. No significant associations were found between sleep quality and the genotypic and allelic distributions of the Per3 gene SNP's in the BD sample.
The present study has limitations that should be well thought out when interpreting the results. Small sample size would be a major limitation and future studies should make efforts in achieving bigger samples of BD patients.
Several hypotheses can be thrown to try to understand the HWE deviations found in genetic association studies. Genotyping errors are one of the most common causes of deviation from HWE, usually resulting in increased number of heterozygotes from the expected.3839 In the other hand, when the number of heterozygotes are below the expected, one of the most likely possibilities to explain a HWE deviation is due to ethnic stratification and selection of the samples.383940 In our study, the HWE deviation of the SNP rs10462020 was due to a deficiency of heterozygotes than would be predicted by the HWE law.
The population stratification arises from differences between distinct ethnic groups.40 In addition, the Brazilian population presents considerable degree of miscegenation,414243 which could contribute to the HWE deviation found. Other hypotheses have also been thrown while trying to explain the HWE deviations eventually found in genetic studies.40 Some authors suggest that the deviation from HWE may be exactly an evidence of a genetic association.40 In other words, the deviation itself could accurately represent a possible role of the polymorphism in the predisposition to the disease assessed.444546
It has been suggested that the genetic vulnerability of BD would be of complex polygenic nature, with multiple susceptibility genes each conferring small effects.47 Therefore, the individual effect of each gene in determining the vulnerability to BD would be of minor magnitude and may not always reach statistical significance.47 However, in addition to the effect of each individual gene, other genetic factors may confer increased risk for BD, including structural genomic variations (CNV's), mitochondrial DNA variation and genetic and epigenetic interactions.47 In fact, some recent publication have demonstrated associations of epigenetic interactions between specific circadian genes, including the Per3 gene, and BD.4849 Thus, at least partially, the past mixed results of association between the Per3 gene and BD may be related to all these referred reasons as well as to the different compositions of the samples of each study, as mentioned in the last paragraph.
Still as regards to the mixed results of association between BD and the Per3 gene, previous studies showed that the regulation of the circadian system would be far more complex than initially proposed.5051 Consequently, in addition to the core clock genes directly involved in the central regulation of the circadian system (Clock, Artnl1, Cry1, Cry2, Per1, Per2, and Per3), new evidence suggest that hundreds of genes may play important role for the intricate process of the circadian regulation system.52 As a result, changes accumulated in a numerically larger set of circadian genes may possibly work in an integrated way towards an underlying vulnerability factor for BD.525354
In this sense, the use of new approaches in order to pool into the same analysis some of the genetic factors related to the vulnerability of BD can lead to advances in understanding the underlying susceptibility of the disorder.53 Convergent functional genomics and pathway analysis studies try to overcome this difficulty by integrating biological findings from various sources to genetic results arising from GWAS studies.525354 Therefore, instead of analyzing the association of a gene individually, which may frequently not reach the statistical significance threshold, these studies evaluate groups of genes involved in the same physiological pathway functionally related to the studied disease. Recently, some such studies that evaluated several genes involved in the regulation of the circadian system found positive association with BD.52535455 Thus, it is possible that these new methodologies will be able to better translate the polygenic nature of the genetic vulnerability to BD.
In addition to the limitations of association case-control studies involving a single candidate gene in identifying the underlying heritability of complex genetic disease such as BD, listed in the last paragraphs, some other limitations should be considered when evaluating the results of this study. The PSQI assesses sleep quality and its use is widespread both in the clinical setting and in scientific research.32 However, a major limitation of this study involves the low PSQI's ability to capture, in its full complexity, the various aspects of the socalled phenomenon of sleep-wake cycle. In fact, researchers claim sleep as one of the least understood topics in biological sciences.8 Furthermore, while evaluating sleep with the PSQI in the period related to the last 30 days, one cannot exclude the interference of a recall bias. Moreover, by capturing an average behavior, the instrument does not gather information regarding the variability of the sleep patterns day after day, which would be best captured by actigraphy or sleep diaries. Therefore, other phenomena related to the sleep function, which may be not fully captured by the PSQI, might be associated with the Per3 gene. However, despite sleep questionnaires are considered subjective sleep parameters, some evidence showed correlation of these with objective parameters measured by actigraphy and polysomnography in a sample from patients with BD.56
One last relevant issue that needs to be further addressed is the possible shared vulnerability of Sleep Disorders and BD. In fact, since sleep was recognized to be controlled genetically, researchers have showed that the mechanisms of sleep regulation and pathways, such as the circadian system and some neurotransmitter signaling, have conserved effects on sleep across a wide range of species up to humans.8 At the same time, genome-wide association studies (GWAS) have exposed a few loci rising vulnerability to primary Sleep Disorders as narcolepsy and restless leg syndrome.8 Thus, it is likely that, at least partially, Sleep Disorders and BD may join common liability factors.
Some positive aspects of this report should be highlighted. This is, to the best of our knowledge, the first report finding direct and significant association of the Per3 gene polymorphisms and BD in Brazilian and South American samples, as well, of BD patients. This should be of some relevance since Brazilian population has a large degree of miscegenation, as mentioned before, if compared with Caucasian populations. A previous study assessing a Brazilian sample of BD had found association of poor sleep quality and the Per3 gene, but not with BD itself.57 Another interesting aspect of our study is that we found evidence of association of BD with polymorphisms of a gene (Per3) considered a member of the core “clock” circadian system regulation. Additionally, the control group of our report did not differ significantly from cases (BD group) in clinical variables such as age and gender. This is important because researchers claim that one of the main causes of type 1 error in genetic association studies refers to issues regarding specific aspects of the control groups, such as poorly matched control groups.
In this study, we demonstrated evidences of genetic association between the Per3 gene and BD. Our results are consistent with recent findings showing association of the Per3 gene polymorphisms with improvement in depressive symptoms,58 with a family history of suicidal behavior,59 and with sleep disturbances in different samples of BD patients.48 Therefore, even taking into account the limitations of the current study, the results of association between the Per3 gene and BD in our sample may bring additional strength to the former evidences of association among the Per3 gene and BD. Furthermore, this finding is consistent with previous results of association of BD with other genes directly or indirectly related to the circadian regulation.60 Still, if considered together with all the scope of evidences implicating sleep and circadian disturbances with the fundamental pathophysiology of BD,59 as reinforced by recent results of convergent functional genomics and pathway analysis studies,52535455 our findings may enhance the core rationale connecting circadian regulation abnormalities to the neurobiology of BD.
In upcoming studies, assessing multiple control samples would minimize bias due to population stratification. Another interesting approach would be the use of prospective control samples, since generally these control groups are built retrospectively. Furthermore, given the multifactorial nature of BD, studies and techniques that provide the convergence of genetic association, gene expression and epigenetic data should be encouraged. Likewise, future efforts of researchers might focus in testing the shared impact of sets of genes related to the circadian regulation instead of testing one or a few genes individually, since it is reasonable to hypothesize that, according to the complex polygenic nature of BD, the underlying predisposition may be related to the disrupted function of several circadian genes which would, in turn, translate a disrupted circadian system regulation in BD.
Acknowledgments
This study was supported by Instituto Nacional de Ciência e Tecnologia de Medicina Molecular (INCT/CNPq/MCT, FAPEMIG), CNPq Grant No. 473674/2009-2, and FAPEMIG Grant PPM-IV.
References
2. Rivkees SA, Reppert SM. Perinatal development of day-night rhythms in humans. Horm Res 1992;37(Suppl 3):99-104. PMID: 1427650.


3. Reppert SM, Weaver DR. Coordination of circadian timing in mammals. Nature 2002;418:935-941. PMID: 12198538.


5. Miller JD, Morin LP, Schwartz WJ, Moore RY. New insights into the mammalian circadian clock. Sleep 1996;19:641-667. PMID: 8958635.


6. Ko CH, Takahashi JS. Molecular components of the mammalian circadian clock. Hum Mol Genet 2006;15:R271-R277. PMID: 16987893.


7. Hastings M, O’Neill SJ, Maywood ES. Circadian clocks: regulators of endocrine and metabolic rhythms. J Endocrinol 2007;195:187-198. PMID: 17951531.


8. Sehgal A, Mignot E. Genetics of sleep and sleep disorders. Cell 2011;146:194-207. PMID: 21784243.



9. Coomans CP, Lucassen EA, Kooijman S, Fifel K, Deboer T, Rensen PC, et al. Plasticity of circadian clocks and consequences for metabolism. Diabetes Obes Metab 2015;17(Suppl 1):65-75. PMID: 26332970.


10. Liu C, Chung M. Genetics and epigenetics of circadian rhythms and their potential roles in neuropsychiatric disorders. Neurosci Bull 2015;31:141-159. PMID: 25652815.



11. Atkinson M, Kripke DF, Wolf SR. Autorhytmometry in manic-depressives. Chronobiologia 1975;2:325-335. PMID: 1222678.

12. Kripke DF, Mullaney DJ, Atkinson M, Wolf S. Circadian rhythm disorders in manic-depressives. Biol Psychiatry 1978;13:335-351. PMID: 667233.

13. Cajochen C, Krauchi K, Wirz-Justice A. Role of melatonin in the regulation of human circadian rhythms and sleep. J Neuroendocrinol 2003;15:432-437. PMID: 12622846.


14. Soreca I. Circadian rhythms and sleep in bipolar disorder: implications for pathophysiology and treatment. Curr Opin Psychiatry 2014;27:467-471. PMID: 25211500.


15. Rocha PMB, Neves FS, Correa H. Significant sleep disturbances in euthymic bipolar patients. Comp Psychiatry 2013;54:1003-1008.

16. Kerner B. Toward a deeper understanding of the genetics of bipolar disorder. Front Psychiatry 2015;6:105PMID: 26283973.



17. Craddock N, Sklar P. Genetics of bipolar disorder: successful start to a long journey. Trends Genet 2009;25:99-105. PMID: 19144440.


18. Etain B, Milhiet V, Bellivier F, Leboyer M. Genetics of circadian rhythms and mood spectrum disorders. Eur Neuropsychopharmacol 2011;21(Suppl 4):S676-S682. PMID: 21835597.


19. Zylka MJ, Shearman LP, Weaver DR, Reppert SM. Three period homologs in mammals: differential light responses in the suprachiasmatic circadian clock and oscillating transcripts outside of brain. Neuron 1998;20:1103-1110. PMID: 9655499.


20. Osland TM, Ferno J, Havik B, Heuch I, Ruoff P, Laerum OD, et al. Lithium differentially affects clock gene expression in serum-shocked NIH-3T3 cells. J Psychopharmacol 2011;25:924-933. PMID: 20837565.


21. Cichon S, Schumacher J, Muller DJ, Hurter M, Windemuth C, Strauch K, et al. A genome screen for genes predisposing to bipolar affective disorder detects a new susceptibility locus on 8q. Hum Mol Genet 2001;10:2933-2944. PMID: 11741836.


22. Curtis D, Kalsi G, Brynjolfsson J, McInnis M, O’Neill J, Smyth C, et al. Genome scan of pedigrees multiply affected with bipolar disorder provides further support for the presence of a susceptibility locus on chromosome 12q23-q24, and suggests the presence of additional loci on 1p and 1q. Psychiatr Genet 2003;13:77-84. PMID: 12782963.


23. Nievergelt CM, Kripke DF, Barrett TB, Burg E, Remick RA, Sadovnick AD, et al. Suggestive evidence for association of the circadian genes PERIOD3 and ARNTL with bipolar disorder. Am J Med Genet B Neuropsychiatr Genet 2006;141B:234-241. PMID: 16528748.



24. Mansour HA, Wood J, Loque T, Chowdari KV, Dayal M, Monk TH, et al. Association study of eight circadian genes with bipolar I disorder, schizoaffective disorder and schizophrenia. Genes Brain Behav 2006;5:150-157. PMID: 16507006.


25. Benedetti F, Dallaspezia S, Colombo C, Pirovano A, Marino E, Smeraldi E. A length polymorphism in the circadian clock gene Per3 influences age at onset of bipolar disorder. Neurosci Lett 2008;445:184-187. PMID: 18789374.


26. Kripke DF, Nievergelt CM, Joo E, Shekhtman T, Kelsoe JR. Circadian polymorphisms associated with affective disorders. J Circadian Rhythms 2009;7:2PMID: 19166596.



27. Soria V, Martinez-Amoros E, Escaramis G, Valero J, Peres-Egea R, Garcia C, et al. Differential association of circadian genes with mood disorders: CRY1 and NPAS2 are associated with unipolar major depression and CLOCK and VIP with bipolar disorder. Neuropsychopharmacology 2010;35:1279-1289. PMID: 20072116.



28. Dallaspezia S, Lorenzi C, Pirovano A, Colombo C, Smeraldi E, Benedetti F. Circadian clock gene Per3 variants influence the postpartum onset of bipolar disorder. Eur Psychiatry 2011;26:138-140. PMID: 21316201.


29. Mansour HA, Talkowski ME, Wood J, Chowdari KV, McClain L, Prasad K, et al. Association study of 21 circadian genes with bipolar I disorder, schizoaffective disorder, and schizophrenia. Bipolar Disord 2009;11:701-710. PMID: 19839995.



30. Karthikeyan R, Marimuthu G, Ramasubramanian C, Arunachal G, BaHammam AS, Spence DW. Association of Per3 length polymorphism with bipolar I disorder and schizophrenia. Neuropsychiatr Dis Treat 2014;10:2325-2330. PMID: 25525361.


31. Buysse DJ, Reynolds CF 3rd, Monk TH, Berman SR, Kupfer DJ. The Pittsburgh Sleep Quality Index: a new instrument for psychiatric practice and research. Psychiatry Res 1989;28:193-213. PMID: 2748771.


32. Bertolazi AN, Fagondes SC, Hoff LS, Dartora EG, Miozzo IC, de Barba ME, et al. Validation of the Brazilian Portuguese version of the Pittsburgh Sleep Quality Index. Sleep Med 2011;12:70-75. PMID: 21145786.


33. American Psychiatry Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-IV-TR). 4th Ed. Washington DC: APA; 2000.
34. Hamilton M. A rating scale for depression. J Neurol Neurosurg Psychiatry 1960;23:56-62. PMID: 14399272.



35. Young RC, Biggs JT, Ziegler VE, Meyer DA. A rating scale for mania: reliability, validity, and sensitivity. Br J Psychiatry 1978;133:429-433. PMID: 728692.


36. Amorim P. Mini International Neuropsychiatric Interview (MINI): validation of a brief interview for the diagnosis of mental disorders. Rev Bras Psiquiatr 2000;22:106-115.

37. Lahiri DK, Nurnberger JI Jr. A rapid non-enzymatic method for the preparation of HMW DNA from blood for RFLP studies. Nucleic Acids Res 1991;19:5444PMID: 1681511.



38. Zintzaras E. Impact of Hardy-Weinberg equilibrium deviation on allele-based risk effect of genetic association studies and meta-analysis. Eur J Epidemiol 2010;25:553-560. PMID: 20526652.


39. Maynard Smith J. Evolutionary Genetics. . New York: Oxford University Press; 1998.
40. Wittke-Thompson JK, Pluzhinikov A, Cox NJ. Rational inferences about departures from Hardy-Weinberg equilibrium. Am J Hum Genet 2005;76:967-986. PMID: 15834813.



41. Fausto B. História do Brasil. 14th Ed. São Paulo: EDUSP; 2012.
42. Salzano FM, Bortolini MC. The Evolution and Genetics of Latin American Population. Cambridge: Cambridge University Press; 2002.
43. Santos RV, da Silva GO, Gibbon S. Pharmacogenomics, human genetic diversity and the incorporation and rejection of color/race in Brazil. Biosocieties 2015;10:48-69. PMID: 26290677.



44. Balding DJ. A tutorial on statistical methods for population association studies. Nat Rev Genet 2006;7:781-791. PMID: 16983374.


45. Wang J, Shete S. Testing departure from Hardy-Weinberg proportions. Methods Mol Biol 2012;850:77-102. PMID: 22307695.


46. Nielsen DM, Ehm MG, Weir BS. Detecting marker-disease association by testing for Hardy-Weinberg disequilibrium at a marker locus. Am J Hum Genet 1998;63:1531-1540. PMID: 9867708.



48. Dmitrzak-Weglarz MP, Pawlak JM, Maciukiewicz M, Moczko J, Wilkosc M, Leszcynska-Rodziewics A, et al. Clock gene variants differentiate mood disorders. Mol Biol Rep 2015;42:277-288. PMID: 25258123.


49. Maciukiewicz M, Dmitrzak-Weglarz M, Pawlak J, LeszczynskaRodziewiecz , Zaremba D, Skibinska M, et al. Analysis of genetic association and epistasis interactions between circadian clock genes and symptom dimensions of bipolar affective disorder. Chronobiol Int 2014;31:770-778. PMID: 24673294.


50. Ueda HR, Hayashi S, Chen W, Sano M, Shigeyoshi Y, Lino M, et al. System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nature Genet 2005;37:187-192. PMID: 15665827.


51. Zhang EE, Liu AC, Hirota T, Miraglia LJ, Welch G, Ponqsawakul PY, et al. A genome wide RNAi screen for modifiers of the circadian clock in human cells. Cell 2009;139:199-210. PMID: 19765810.



52. McCarthy MJ, Nievergelt CM, Kelsoe JR, Welsh DK. A survey of genomic studies supports association of circadian clock genes with bipolar disorder spectrum illnesses and lithium response. PLoS One 2012;7:e32091. PMID: 22384149.



53. Nurnberger JI Jr. Genetics of bipolar disorder: where we are and where we are going. Depress Anxiety 2012;29:991-993. PMID: 23208953.


54. Patel SD, Le-Niculescu H, Koller DL, Green SD, Lahiri DK, McMahon FJ, et al. Coming to grips with complex disorders: genetic risk prediction in bipolar disorder using panels of genes identified through convergent functional genomics. Am J Med Genet B Neuropsychiatr Genet 2010;153B:850-877. PMID: 20468069.


55. Le-Niculescu H, Patel SD, Bhat M, Kuczenski R, Faraone SV, Tsuang MT, et al. Convergent functional genomics of genome-wide association data for bipolar disorder: comprehensive identification of candidate genes, pathways and mechanisms. Am J Med Genet B Neuropsychiatr Genet 2009;150B:155-181. PMID: 19025758.


56. Kaplan KA, Talbot LS, Gruber J, Harvey AG. Evaluating sleep in bipolar disorder: comparison between actigraphy, polysomnography, and sleep diary. Bipolar Disord 2012;14:870-879. PMID: 23167935.



57. Rocha PM, Neves FS, Alvarenga NB, Hughet RB, Barbosa IG, Correa H. Association of Per3 gene with bipolar disorder: comment on “Association study of 21 circadian genes with bipolar I disorder, schizoaffective disorder and schizophrenia”. Bipolar Disord 2010;12:875-876. PMID: 21176035.


58. Dallaspezia S, Locatelli C, Lorenzi C, Pirovano A, Colombo C, Benedetti F. Sleep homeostatic pressure and PER3 VNTR gene polymorphism influence antidepressant response to sleep deprivation in bipolar depression. J Affect Disord 2016;192:64-69. PMID: 26707349.


59. Pawlak J, Dmitrzak-Weglarz M, Maciukiewicz M, Wilkosc M, Leszcznska-Rodziewicz A, Zaremba D, et al. Suicidal behavior in the context of disrupted rhythmicity in bipolar disorder--data from an association study of suicide attempts with clock genes. Psychiatry Res 2015;226:517-520. PMID: 25724486.


60. Milhiet V, Boudebesse C, Bellivier F, Drouot X, Henry C, Leboyer M, et al. Circadian abnormalities as markers of susceptibility in bipolar disorders. Front Biosci (Schol Ed) 2014;6:120-137. PMID: 24389266.