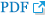|
 |
- Search
| Psychiatry Investig > Volume 15(5); 2018 > Article |
|
Abstract
Objective
The aim of this study was to examine the change in the hippocampal volume and shape in early-onset mild cognitive impairment (EO-MCI) associated with the APOE ε4 carrier state.
Methods
This study had 50 subjects aged 55-63 years, all of whom were diagnosed with MCI at baseline via the Korean version of the Consortium to Establish a Registry for Alzheimer’s Disease Assessment Packet. The EO-MCI patients were divided into the MCI continued (MCIcont) and Alzheimer’s disease (AD) converted (ADconv) groups 2 years later. The hippocampal volume and shape were measured for all the subjects. The local shape analysis (LSA) was used to conduct based on the 2-year-interval magnetic resonance imaging scans.
Results
There was a significant correlation between APOE ε4 allele and hippocampal volume atrophy. Over two years, the volume reduction in the left hippocampus was found to be faster than that in the right hippocampus, especially in the APOE ε4 carriers. LSA showed that the 2 subfields were significantly affected in the left hippocampus.
Mild cognitive impairment (MCI), especially the amnestic type of MCI, refers to a prodromal stage of Alzheimer’s disease (AD) during which patients display cognitive impairment without apparent disabilities in their activities of daily living. Patients with MCI show memory dysfunction, and 10-20% of MCI annually progresses into AD [1,2]. Atrophy of the hippocampus is one of the most sensitive biological indicators of AD, and hippocampal volumetry is the best-established structural biomarker for AD, especially for early diagnosis [3-8]. Earlier studies have reported that hippocampal atrophy can to some extent predict which patients with MCI will subsequently convert to AD [9,10], and that the AD converters (ADconv) group had an enhanced rate of hippocampal volume loss [11]. The ADconv group had a significantly higher rate of hippocampal volume loss compared to the MCI continued (MCIcont) group [12,13].
There are two main subtypes of AD: early-onset AD (EO-AD) and sporadic AD. A very small percentage of the people who develop AD have the EO variety, which is classified as the beginning of the disease presentation before the age of 65. Researchers have discovered three genes in which mutations cause EO-AD [14]. These known genetic mutations, however, account for only 2% of all the cases of EO-AD [15]. The other group, sporadic AD, is most commonly termed “late-onset AD (LO-AD).” It is defined by the disease presentation after the age of 65, and it is well known that the apolipoprotein E (APOE) ε4 allele can influence it. According to the current viewpoint, however, classifying AD into EO or LO is probably not useful from a mechanistic point of view because mutations in the 3 genes can be found in both EO and LO. Similarly, APOE ε4 allele increases the risk of AD in both EO-AD and LO-AD. Accordingly, EO-AD that does not have an autosomal dominant pattern can share similar characteristics with LO-AD and may be influenced by the APOE ε4 allele, similar to LO-AD [14,16]. Considering that MCI is a transitional state between normal aging and AD, it may be speculated that MCI can also be divided into EO-MCI and LO-MCI according to the age of symptom onset. EO-MCI and LO-MCI may be expected to have patterns of differences similar to those between EO-AD and LO-AD because EO-AD and LO-AD can have different clinical features and anatomical substrates [17].
The identification and characterization of patients with EOMCI may have clinical significance because the potential impact of cognitive impairment on vocational and social activities is probably greater in the young than in the older MCI patients [18]. To the authors’ knowledge, however, there has been no study exploring EO-MCI and comparing it with LO-MCI in terms of the imaging characteristics.
Recent research has shown that the APOE ε4 allele is well known as the risk factor not only for LO-AD but also for EOAD [14,16,19]. Studies involving EO-AD subjects under 65 years associated with the APOE ε4 allele are not common [17], and the research on EO-MCI is still limited because all the studies so far have focused on LO-MCI/AD, which is more common than EO-MCI/AD [20]. Additionally, the hippocampus is not a homogeneous structure but consists of several subfields with distinct histological characteristics: the subiculum, the three cornu ammonis (CA) sectors (CA1, CA2, and CA3), and the dentate gyrus [21]. Therefore, this neuroimaging study is expected to explain some of the characteristics of the MCIcont and ADconv groups in terms of volume- and shape- based morphometrics in the EO-MCI subjects using hippocampal volumetry and local shape analysis (LSA) [7,22]. The authors targeted EO-MCI subjects under 65 years who had very important diagnostic values, divided them into two groups according to their APOE ε4 status, and checked their brain magnetic resonance imaging (MRI) scans at baseline and 2 year later. This study was done to elucidate the changes in the hippocampal volume and shape in the EO-MCI subjects according to their APOE ε4 status and according to their AD dementia conversion.
The study subjects were selected while in the implementation of the dementia early screening project of the Ministry of Health and Welfare from March 2008 to February 2016. A skilled geropsychiatrist and trained research nurses performed examinations with the Korean version of the Consortium to Establish a Registry for Alzheimer’s Disease Assessment Packet (CERAD-K) for the elderly aged 55-63 years who were residing in the Chungbuk area, and those who were finally diagnosed with amnestic MCI at baseline were targeted for the study. Of them, those who had a Clinical Dementia Rating (CDR) score of 0.5 in the examination, who did not have a history of cerebrovascular disease, who did not have a major depressive disorder based on Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition, Text Revision (DSM-IV-TR), and who had a Modified Hachinski Ischemic Scale [23] score of 3 or less were included in the study. On the other hand, those with delirium, confusion, or other clouded consciousness; those with neurological diseases such as Parkinson’s disease, stroke, tumors, and normal pressure hydrocephalus; those with evidence of severe cerebrovascular disease; and those who had medical conditions that might have serious effects on their cognitive functions or who were taking drugs affecting their cognitive functions were excluded. A total of 66 persons at baseline were selected as study subjects and gave their consent for genetic testing to check the presence of the APOE ε4 allele. With the subjects who did not undergo genetic testing being excluded, a total of 53 study subjects were selected. All the study subjects went through brain MRI scan at baseline and 2 years later. As a result, based on the brain MRI scan, a total of 50 subjects were included in this study: 35 subjects who met the aforementioned diagnosis criteria and were diagnosed with MCI (MCIcont), and 15 subjects who were diagnosed with AD dementia (ADconv). Meanwhile, three subjects were excluded. Of these 3 subjects, two subjects were excluded due to vascular dementia, and one subject was excluded due to frontotemporal dementia. Written informed consent was obtained from each patient, and this study was reviewed and approved by the local IRB (KUCH 2017-12-033).
Brain MRI scan was conducted at the Department of Radiology of a university hospital, using a 1.5-Tesla scanner (MRT200PP3, Toshiba, Japan). With the following parameters, the MRI scans were obtained: repetition time (TR)=15 ms, echo time (TE)=5 ms, field of view (FOV)=240×240 mm, flip angle=20°, slice thickness=1.5 mm, matrix size=256×256, acquisition time=3.06 min, resolution=0.93×0.93×1.5 mm, yielding 3D T1-weighted images.
The hippocampus was manually labeled by a rater in the contiguous coronal brain sections using BrainSuite24 and guided by well-established protocols (http://users.loni.ucla.edu/~narr/protocol.php?q=hippotrace). The same rater repeated this process three times. After manually labeling the hippocampus as described above, the global left and right hippocampal volumes were established (in mm3) based on the number of voxels constituting the hippocampus labels.
To compare the baseline hippocampal volume (i.e., hippocampal volume at the time of the initial MRI scan) of the study groups, the hippocampal volume measurements were normalized for inter-individual variation in head size, age, and gender [25]. The hippocampal volumes in each individual were summed and divided by the total intracranial volume (ICV) of that particular patient to control for variation in head size [12]. ICV measurement was done automatically using the Lab of NeuroImaging (LONI) pipeline system of University of Southern California (USC) [26]. The hippocampal atrophy rates were calculated based on the reduction percentages of the hippocampal volumes between the 2-year-interval MRI scans.
The LSA pipeline workflow was used to conduct local (pervertex) post-hoc statistical analyses of the shape differences between the two-year-interval brain MRI scans in the left and right hippocampus. In the LSA pipeline workflow, the 3D structural MRI data are first preprocessed (skull-stripped, spatially normalized, parcellated) [26,27], then shape models of the hipocampi are generated as genus-zero two-dimensional manifolds [28,29]. By traversing the triangulated boundary manifolds (per-vertex), statistical significance maps representing the group differences in two complementary shape metrics are obtained. The radial distance measures at each vertex encode the magnitude and direction of the local shape morphometry, which quantify the discrepancy between each subject as the “mean shape” (boundary) for both hippocampi. Probability values corresponding to the test statistics are overlaid on the mean boundary shape for each region to illustrate the group differences.
Group differences between the MCIcont and ADconv groups were analyzed by Student’s t-test for continuous variables such as age or education years or χ2 (chi-squared) test for categorical variables such as the gender and the APOE ε4 carrier status. According to the presence of ε4, both hippocampi volumes were analyzed with a 2-year interval via repeated-measures ANOVA (analysis of variance), controlling the age and gender, which might affect the results. In addition, for group comparisons, the unpaired t-test (two-tailed) was used together with Mann-Whitney U test as a non-parametric statistical analysis. Effects of the age, gender, and APOE ε4 allele on the atrophy rate of both hippocampi were measured using general linear model (GLM) analyses for the MCIcont and ADconv groups respectively. For the analysis, SPSS version 20.0 for the Windows program was used.
Moreover, the statistical method of the multiple linear regression (MLR) model was used to conduct the LSA analysis, and a design matrix that included age, gender, and APOE ε4 status as regressors was used. The EO-MCI subjects were analyzed as a whole and by group (MCIcont and ADconv groups), using the anatomical morphometric measures as predictors of the diagnosis as the response variables. For LSA to analyze the shape, the MLR model was applied using the Laboratory of NeuroImaging (LONI) pipeline system of USC.
Among the total of 50 study subjects, 35 (70%) were included in MCIcont group and 15 (30%) were included in the ADconv group 2 years later. The mean age was 63.81±3.19 (M±SD) years in the MCIcont group and 63.64±3.01 (M±SD) years in the ADconv group, not showing a significant difference (t=0.149, p=0.883). The mean education years was not showing a significant difference (t=0.276, p=0.785). The MMSEKC score was 25.25±1.43 (M±SD) in the MCIcont group and 20.24±3.12 (M±SD) in the ADconv group, showing a significant difference (t=1.134, p=0.001). The APOE ε4 status was not showing a significant difference (χ2=0.736, p=0.391). The demographic and clinical data are described in Table 1.
As a result of the re-measurement of the hippocampal volume with a 2-year interval from the initial measurement at the baseline, there was a significant change in the volume regardless of the presence of APOE ε4 in both the MCIcont and ADconv groups. In the case of the left hippocampus, a more statistically significant result was shown in the ADconv group especially when APOE ε4 allele was present (F=11.470, p=0.001) (Table 2).
In the case of the left hippocampus, the atrophy rate was significantly higher in the ADconv group than in the MCIcont group within the APOE ε4 carrier group (t=-2.91, p=0.005, tied p=0.003). Even within the ADconv group, the atrophy rate was significantly higher in the APOE ε4 carrier group than in the APOE ε4 non-carrier group (t=-2.58, p=0.016, tied p=0.010). In the case of the right hippocampus, the atrophy rate tended to be higher but it was not statistically significant (Table 3).
In the MCIcont group, the atrophy rate of the left hippocampus was higher in the APOE ε4 carrier group in the GLM analysis, regardless of the age and gender (F=5.532, p=0.025). The atrophy rate of the right hippocampus tended to be higher in the APOE ε4 carrier group, but it was not statistically significant (F=3.82, p=0.056) (Table 4).
In the ADconv group, the atrophy rate of the left hippocampus was higher in the APOE ε4 carrier group in the GLM analysis, regardless of the age and gender (F=12.978, p=0.001). Even the atrophy rate of the right hippocampus was still significantly higher in the APOE ε4 carrier group (F=11.05, p=0.007) (Table 4).
Both hippocampi were chosen to find out if their shape measures are also associated with the changes in the 2-year-interval MRI scans (regressor). The result of the LSA of the hippocampus in the MRI scanning with a 2-year interval showed no significant outcome surviving the false discovery rate (FDR) correction in the analysis of the difference between the MCIcont and ADconv groups and in the analysis based on the presence of APOE ε4. Only when the MRI values of the total EO-MCI group were compared without controlling any variable with a 2-year interval was it found that the shape of the left hippocampus was significantly changed in the hippocampal radial distance and survived the FDR correction. In the case of the right hippocampus, however, no case survived the FDR correction. All the results that survived the FDR correction are shown as p-value color maps in Figure 1. Figure illustrates the significant atrophic change in the hippocampal radial distance between the 2-year follow-up scans at the left hippocampus.
The main findings in this study were that the hippocampal atrophy rates predict conversion to dementia in MCI patients, and that the left hippocampal volume associated with the APOE ε4 allele seems to be the best marker for the ADconv group in EO-MCI. In this study, the atrophy rate of the left hippocampus was significantly higher than the right hippocampus in the APOE ε4 carriers of the MCIcont group. The atrophy rate of both hippocampi in the ADconv group was significantly higher in the APOE ε4 carriers than in the APOE ε4 non-carriers, but the atrophy rate of the left hippocampus was higher than the atrophy rate of the right hippocampus. In LSA for hippocampal shape measure, the radial distance atrophy in the left hippocampus was significantly changed regardless of APOE ε4 and the AD conversion status. A previous 3-year clinical follow-up LO study reported that the mean annualized rates of hippocampal atrophy were control, 1.73%; MCIcont, 2.55%; ADconv, 3.69%; and AD, 3.5%. The main findings of the study were that the hippocampal volume predicts conversion to dementia in LO-MCI patients, and that the left hippocampal volume seems to be the best marker for conversion [12]. Another LO study reported that the mean annualized rates of hippocampal atrophy were MCIcont, 3.0% and ADconv, 4.1% [30]. Taken together from the LO studies, including the two aforementioned studies, the volume of the hippocampus is reduced by 1.55-2.09% annually in normal persons in general, while it is reduced by 2.55-3.0% annually in the MCIcont group and 3.69-4.1% annually in the ADconv group [31]. Thus, if the volume of the hippocampus is reduced by more than 3.0% annually in LO-MCI patients, the possibility of developing AD dementia is relatively higher. In this EO study, it was found that the annualized rates of hippocampal atrophy by 2-year follow-up were MCIcont, 2.38% and ADconv, 2.79%. This EO study on hippocampal atrophy rates revealed that the rates were lower than those in the previous LO studies. This difference can be explained from a previous topographic study which indicate that EO-AD and LO-AD differ in their typical topographic patterns of brain atrophy, suggesting different predisposing or etiological factors [19]. Given that EO-AD and LO-AD have different clinical features and anatomical substrates, EO-MCI and LO-MCI may be expected to have patterns of differences similar to those between EO-AD and LO-AD [17]. As one more reason for slow atrophy in EO-MCI, it can be said that EO-MCI is a less characterized group and is probably more heterogeneous than LO-MCI, including more patients with other comorbidities [32,33]. Therefore, the relatively lower prevalence of AD among people under 65 years of age, and the higher prevalence of other types of dementia [18], can lead to slower atrophy in EO-MCI than in LO-MCI [34]. These reports may say that the hippocampal atrophy can be less influenced in EO-MCI/AD than in LO-MCI/AD. It is difficult to conclusively say, however, that the hippocampal volume atrophy is trivial because the atrophy in EO-AD/MCI is slow. EO-MCI/AD may first affect another area (including neocortex) but will affect the hippocampus as well, on the extension of LO-MCI/AD, and its atrophy can be still important in predicting AD dementia conversion. This study showed that the degree of hippocampal atrophy of EO-MCI does not seem to be larger than that of LO-MCI, but it can still be higher associated with the moderating effect of the APOE ε4 allele. Even though the hippocampal atrophy is known to be smaller in EOMCI/AD than in LO-MCI/AD, EO and LO can be considered as one continuum since the age of 65 is an arbitrary cut-off, which does not have a true biological meaning [35,36].
In terms of our data, annual EO-AD conversion rate was 15% (30% for 2 years) and was close to the conversion rate in LO-MCI, which can say that EO-MCI and LO-MCI may lie in a continuum. On the other hand, when it comes to hippocampal atrophy rate, it was slower in EO-MCI than in LO-MCI, which can show us that the atrophy in hippocampus, is less influenced in EO-MCI.
In line with volume-based comparison, our shape-based diffeomorphometric comparison results of LSA revealed that radial distance atrophy in left hippocampus was significantly changed. The radial distance measures the distance from each surface point to the central core of the shape (point of gravitational balance). This metric index captures local expansions or contractions of the developing cortical surface. Previous research shows that the hippocampal formation is unevenly affected by AD pathology, deposits of plaques and tangles being particularly dense in the CA1 field and subiculum [3], and regions corresponding to the CA2/3 fields were remarkably spared [37,38]. Our data are very consistent with the results of this research on hippocampal atrophy rates and associated with previous morphometric comparison research on hippocampus as well.
This study associated with EO-MCI, especially with ADconv group, revealed that faster left hippocampal atrophy in volume especially in APOE ε4 carriers. Similarly, Risacher et al. [39] reported that decreased hippocampal volume (left more than right) was the most robust structural MRI feature. Our study can reveal that left hippocampus is more atrophied in EO-MCI, as a continuum in LO-MCI.
In this study, the moderating effect of the APOE ε4 allele on hippocampal atrophy was shown in both hippocampi atrophy rates, but the moderating effect for the left hippocampus was more remarkable. Several studies reported that the left hippocampus related or not related to the APOE ε4 allele contracts significantly faster in LO-MCI [11,39,40]. This EO-MCI study including LSA for the hippocampus showed the same result. Therefore, volume- and shape- based research should be continuously performed for the issue that the moderating effect of the APOE ε4 allele on the left hippocampus affects the conversion into AD dementia.
There are several limitations to this EO study. First, the sample size was small due to the significant data stratification and lack of available data. EO-AD is rarer than LO-AD, which contributed to the relative weakness of the statistical results. Second, this study did not include neuroimaging data from the LO-MCI subjects and asymptomatic normal controls. Thus, the EO-MCI, LO-MCI, and healthy subjects were not compared. Third, the brain images of EO-AD were not compared with those of LO-AD; thus, the characteristics were cited from other studies, which is also a limitation of this study. Finally, the EO-MCI group can be more heterogeneous than the LO-MCI group, including more patients with other comorbidities. Despite this limitations, the present study is significant in that it is a first attempt to identify the neuroimaging differences between EO-MCI and LO-MCI. Further research with increased number of EO subjects will be needed to identify the neuroimaging characteristics in EO-MCI.
Figure 1.
Left hippocampus radial distance (local shape analysis, FDR corrected). A: Bottom view of left hippocampus. B: Anterosuperior view of left hippocampus. FDR: false discovery rate.
Table 1.
Demographic and clinical characteristics of the subjects
| MCIcont (N=35) | ADConv (N=15) | Statistics t/χ2 | p | |
|---|---|---|---|---|
| Age (years) | 63.81±3.19* | 63.64±3.01 | t=0.149 | 0.883 |
| Education (years) | 9.10±2.55 | 8.91±2.25 | t=0.276 | 0.785 |
| Gender (%) | χ2=0.019 | 0.892 | ||
| Men | 15 (60.0) | 7 (57.2) | ||
| Women | 20 (40.0) | 8 (42.8) | ||
| MMSE-KC | 25.25±1.43 | 20.24±3.12 | t=1.134 | 0.001† |
| APOE ε4 carriers (%) | 10 (25.8) | 6 (40.0) | χ2=0.736 | 0.391 |
Table 2.
Changes in the hippocampal volume between the MCIcont and ADconv groups after 2 years
| Hippocampus |
MCIcont (N=35) |
ADconv (N=15) |
||||||
|---|---|---|---|---|---|---|---|---|
|
APOE ε4 (-) (N=25) |
APOE ε4 (+) (N=9) |
APOE ε4 (-) (N=9) |
APOE ε4 (+) (N=6) |
|||||
| Baseline | 2 years | Baseline | 2 years | Baseline | 2 years | Baseline | 2 years | |
| Left | 2,338±343* | 2,231±344 | 2,318±357 | 2,204±354 | 2,329±364 | 21,94±352 | 2,317±374 | 2,181±363 |
| Right | 2,529±364 | 2,430±367 | 2,519±364 | 2,412±371 | 2,579±239 | 2,468±259 | 2,569±389 | 2,447±387 |
Table 3.
Differences in atrophy rate between the left and right hippocampi (according to the APOE ε4 status)
| Atrophy rate (%/year) |
MCIcont (N=35) |
ADconv (N=15) |
p (2, 4) | p (3, 4) | ||
|---|---|---|---|---|---|---|
| 1; APOEε4 (-) (N=26) | 2; APOEε4 (+) (N=9) | 3; APOEε4 (-) (N=9) | 4; APOEε4 (+) (N=6) | |||
| Left (% of changes) | 2.29±0.89* | 2.46±0.87 | 2.64±1.02 | 2.93±1.18 | 0.005 (t=-2.91)† | 0.016 (t=-2.58)† |
| (tied p=0.003)‡ | (tied p=0.010)‡ | |||||
| Right (% of changes) | 1.96±0.58 | 2.12±0.56 | 2.15±0.63 | 2.34±0.99 | 0.060 (t=-1.97) | 0.054 (t=-2.087) |
| (tied p=0.081)‡ | (tied p=0.064)‡ | |||||
Table 4.
Effect of APOE ε4 on the atrophy rate of both hippocampi in the MCIcont and ADcov groups: results of the GLM analysis
|
MCIcont (N=35) |
ADconv (N=15) |
|||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
Left hippocampus |
Right hippocampus |
Left hippocampus |
Right hippocampus |
|||||||||||||||||
| Type III SS | df | Mean square | F | p | Type III SS | df | Mean square | F | p | Type III SS | df | Mean square | F | p | Type III SS | df | Mean square | F | p | |
| Age | 13.485 | 1 | 13.485 | 3.873 | 0.060 | 30.518 | 1 | 30.518 | 2.583 | 0.118 | 5.271 | 1 | 5.271 | 1.996 | 0.185 | 16.693 | 1 | 16.693 | 3.712 | 0.082 |
| Gender | 3.305 | 1 | 3.305 | 0.476 | 0.612 | 5.625 | 1 | 5.625 | 0.822 | 0.495 | 0.544 | 1 | 0.544 | 0.208 | 0.660 | 0.753 | 1 | 0.753 | 0.167 | 0.690 |
| APOE ε4 | 19.258 | 1 | 19.258 | 5.532 | 0.025* | 45.139 | 1 | 45.139 | 3.82 | 0.059 | 34.281 | 1 | 34.28 | 12.978 | 0.001† | 29.188 | 1 | 29.188 | 11.05 | 0.007† |
REFERENCES
1. Farias ST, Mungas D, Reed BR, Harvey D, DeCarli C. Progression of mild cognitive impairment to dementia in clinic- vs community-based cohorts. Arch Neurol 2009;66:1151-1157.



2. Petersen RC, Roberts RO, Knopman DS, Boeve BF, Geda YE, Ivnik RJ, et al. Mild cognitive impairment: ten years later. Arch Neurol 2009;66:1447-1455.



3. Apostolova LG, Dutton RA, Dinov ID, Hayashi KM, Toga AW, Cummings JL, et al. Conversion of mild cognitive impairment to Alzheimer disease predicted by hippocampal atrophy maps. Arch Neurol 2006;63:693-699.


4. Apostolova LG, Thompson PM, Green AE, Hwang KS, Zoumalan C, Jack CR Jr, et al. 3D comparison of low, intermediate, and advanced hippocampal atrophy in MCI. Hum Brain Mapp 2010;31:786-797.



5. Killiany RJ, Gomez-Isla T, Moss M, Kikinis R, Sandor T, Jolesz F, et al. Use of structural magnetic resonance imaging to predict who will get Alzheimer’s disease. Ann Neurol 2000;47:430-439.


6. Du AT, Schuff N, Amend D, Laakso MP, Hsu YY, Jagust WJ, et al. Magnetic resonance imaging of the entorhinal cortex and hippocampus in mild cognitive impairment and Alzheimer’s disease. J Neurol Neurosurg Psychiatry 2001;71:441-447.



7. Moon SW, Dinov ID, Hobel S, Zamanyan A, Choi YC, Shi R, et al. Structural brain changes in early-onset Alzheimer’s disease subjects using the LONI pipeline environment. J Neuroimaging 2015;25:728-737.



8. Frisoni GB, Fox NC, Jack CR Jr, Scheltens P, Thompson PM. The clinical use of structural MRI in Alzheimer disease. Nat Rev Neurol 2010;6:67-77.



9. Tapiola T, Pennanen C, Tapiola M, Tervo S, Kivipelto M, Hanninen T, et al. MRI of hippocampus and entorhinal cortex in mild cognitive impairment: a follow-up study. Neurobiol Aging 2008;29:31-38.


10. Tanpitukpongse TP, Mazurowski MA, Ikhena J, Petrella JR. Alzheimer’s disease neuroimaging I. Predictive utility of marketed volumetric software tools in subjects at risk for Alzheimer disease: do regions outside the hippocampus matter? AJNR Am J Neuroradiol 2017;38:546-552.



11. Eckerstrom C, Olsson E, Borga M, Ekholm S, Ribbelin S, Rolstad S, et al. Small baseline volume of left hippocampus is associated with subsequent conversion of MCI into dementia: the Goteborg MCI study. J Neurol Sci 2008;272:48-59.


12. Jack CR Jr, Petersen RC, Xu Y, O’Brien PC, Smith GE, Ivnik RJ, et al. Rates of hippocampal atrophy correlate with change in clinical status in aging and AD. Neurology 2000;55:484-489.



13. Jack CR Jr, Shiung MM, Gunter JL, O’Brien PC, Weigand SD, Knopman DS, et al. Comparison of different MRI brain atrophy rate measures with clinical disease progression in AD. Neurology 2004;62:591-600.



14. Ertekin-Taner N. Genetics of Alzheimer disease in the pre- and post-GWAS era. Alzheimers Res Ther 2010;2:3



15. Pericak-Vance MA, Grubber J, Bailey LR, Hedges D, West S, Santoro L, et al. Identification of novel genes in late-onset Alzheimer’s disease. Exp Gerontol 2000;35:1343-1352.


16. Pievani M, Galluzzi S, Thompson PM, Rasser PE, Bonetti M, Frisoni GB. APOE4 is associated with greater atrophy of the hippocampal formation in Alzheimer’s disease. Neuroimage 2011;55:909-919.


17. Ye BS, Seo SW, Lee Y, Kim SY, Choi SH, Lee YM, et al. Neuropsychological performance and conversion to Alzheimer’s disease in early- compared to late-onset amnestic mild cognitive impairment: CREDOS study. Dement Geriatr Cogn Disord 2012;34:156-166.


18. Rossor MN, Fox NC, Mummery CJ, Schott JM, Warren JD. The diagnosis of young-onset dementia. Lancet Neurol 2010;9:793-806.



19. Frisoni GB, Pievani M, Testa C, Sabattoli F, Bresciani L, Bonetti M, et al. The topography of grey matter involvement in early and late onset Alzheimer’s disease. Brain 2007;130:720-730.


20. Padovani A, Gilberti N, Borroni B. The usefulness of biological and neuroimaging markers for the diagnosis of early-onset Alzheimer’s disease. Int J Alzheimers Dis 2011;2011:296374



21. Mueller SG, Schuff N, Raptentsetsang S, Elman J, Weiner MW. Selective effect of Apo e4 on CA3 and dentate in normal aging and Alzheimer’s disease using high resolution MRI at 4 T. Neuroimage 2008;42:42-48.



22. Moon SW, Dinov ID, Kim J, Zamanyan A, Hobel S, Thompson PM, et al. Structural neuroimaging genetics interactions in Alzheimer’s disease. J Alzheimers Dis 2015;48:1051-1063.



23. Rosen WG, Terry RD, Fuld PA, Katzman R, Peck A. Pathological verification of ischemic score in differentiation of dementias. Ann Neurol 1980;7:486-488.


24. Shattuck DW, Leahy RM. BrainSuite: an automated cortical surface identification tool. Med Image Anal 2002;6:129-142.


25. Jack CR Jr, Twomey CK, Zinsmeister AR, Sharbrough FW, Petersen RC, Cascino GD. Anterior temporal lobes and hippocampal formations: normative volumetric measurements from MR images in young adults. Radiology 1989;172:549-554.


26. Dinov I Jr, Lozev K, Petrosyan P, Liu Z, Eggert P, Pierce J, et al. Neuroimaging study designs, computational analyses and data provenance using the LONI pipeline. PLoS One 2010;5.

27. Tu Z, Narr KL, Dollar P, Dinov I, Thompson PM, Toga AW. Brain anatomical structure segmentation by hybrid discriminative/generative models. IEEE Trans Med Imaging 2008;27:495-508.



28. Gutman BA, Hua X, Rajagopalan P, Chou YY, Wang Y, Yanovsky I, et al. Maximizing power to track Alzheimer’s disease and MCI progression by LDA-based weighting of longitudinal ventricular surface features. Neuroimage 2013;70:386-401.



29. Wang Y, Song Y, Rajagopalan P, An T, Liu K, Chou YY, et al. Surfacebased TBM boosts power to detect disease effects on the brain: an N=804 ADNI study. Neuroimage 2011;56:1993-2010.



30. Henneman WJ, Sluimer JD, Barnes J, van der Flier WM, Sluimer IC, Fox NC, et al. Hippocampal atrophy rates in Alzheimer disease: added value over whole brain volume measures. Neurology 2009;72:999-1007.



31. Jack CR Jr, Petersen RC, Xu Y, O’Brien PC, Smith GE, Ivnik RJ, et al. Rate of medial temporal lobe atrophy in typical aging and Alzheimer’s disease. Neurology 1998;51:993-999.



32. Apostolova LG, Di LJ, Duffy EL, Brook J, Elashoff D, Tseng CH, et al. Risk factors for behavioral abnormalities in mild cognitive impairment and mild Alzheimer’s disease. Dement Geriatr Cogn Disord 2014;37:315-326.



33. Sachs-Ericsson N, Moxley JH, Corsentino E, Rushing NC, Sheffler J, Selby EA, et al. Melancholia in later life: late and early onset differences in presentation, course, and dementia risk. Int J Geriatr Psychiatry 2014;29:943-951.


34. Anstey KJ, Cherbuin N, Eramudugolla R, Sargent-Cox K, Easteal S, Kumar R, et al. Characterizing mild cognitive disorders in the young-old over 8 years: prevalence, estimated incidence, stability of diagnosis, and impact on IADLs. Alzheimers Dement 2013;9:640-648.


35. Harvey RJ, Skelton-Robinson M, Rossor MN. The prevalence and causes of dementia in people under the age of 65 years. J Neurol Neurosurg Psychiatry 2003;74:1206-1209.



36. Morris JC. Early-stage and preclinical Alzheimer disease. Alzheimer Dis Assoc Disord 2005;19:163-165.


37. Frisoni GB, Sabattoli F, Lee AD, Dutton RA, Toga AW, Thompson PM. In vivo neuropathology of the hippocampal formation in AD: a radial mapping MR-based study. Neuroimage 2006;32:104-110.


38. Frisoni GB, Ganzola R, Canu E, Rub U, Pizzini FB, Alessandrini F, et al. Mapping local hippocampal changes in Alzheimer’s disease and normal ageing with MRI at 3 Tesla. Brain 2008;131:3266-3276.